Gene Synthesis from Oligo Pools on the Ginkgo’s Reconfigurable Automation Cart (RAC) Platform

Authors: Bradley Barber, Ph.D., Elizabeth Gendreau, Alicia Byrn, Alycia Wong
Gene synthesis - the assembly of short DNA oligonucleotides into full, gene-sized fragments (up to ~3 kb) - is a key technique that enables rapid prototyping of in silico-designed, custom DNA constructs. It bypasses the reliance on existing DNA templates and typically requires less time than classical molecular cloning. Researchers use gene synthesis to evaluate DNA sequence variants, tune protein expression or introduce cis-regulatory elements with precision, supporting applications ranging from therapeutic development to metabolic engineering1-3.
This technical note describes a gene synthesis workflow on Ginkgo’s Reconfigurable Automation Cart (RAC) platform, supporting Ginkgo’s downstream high-throughput cell-free protein synthesis screening pipeline.
The fully automated gene synthesis workflow is cheap and scalable.
We use highly multiplexed DNA oligo pools as the starting material4 and the polymerase chain reaction (PCR) to “fish out” target DNA oligos that are required to make particular gene-length fragments. Through a process analogous to Golden Gate assembly5, we then assemble the gene-length fragments, clone them and verify their DNA sequence.
Automated Biological Workflow
The automated workflow consisted of the following main protocol steps:
- Multiplexed PCR: PCR reactions were set up using the 384-well format Bravo liquid handling RAC, and DNA primers specific to pre-defined sub-pools. The Automated Thermal Cycling (ATC) RAC was used to perform thermal cycling.
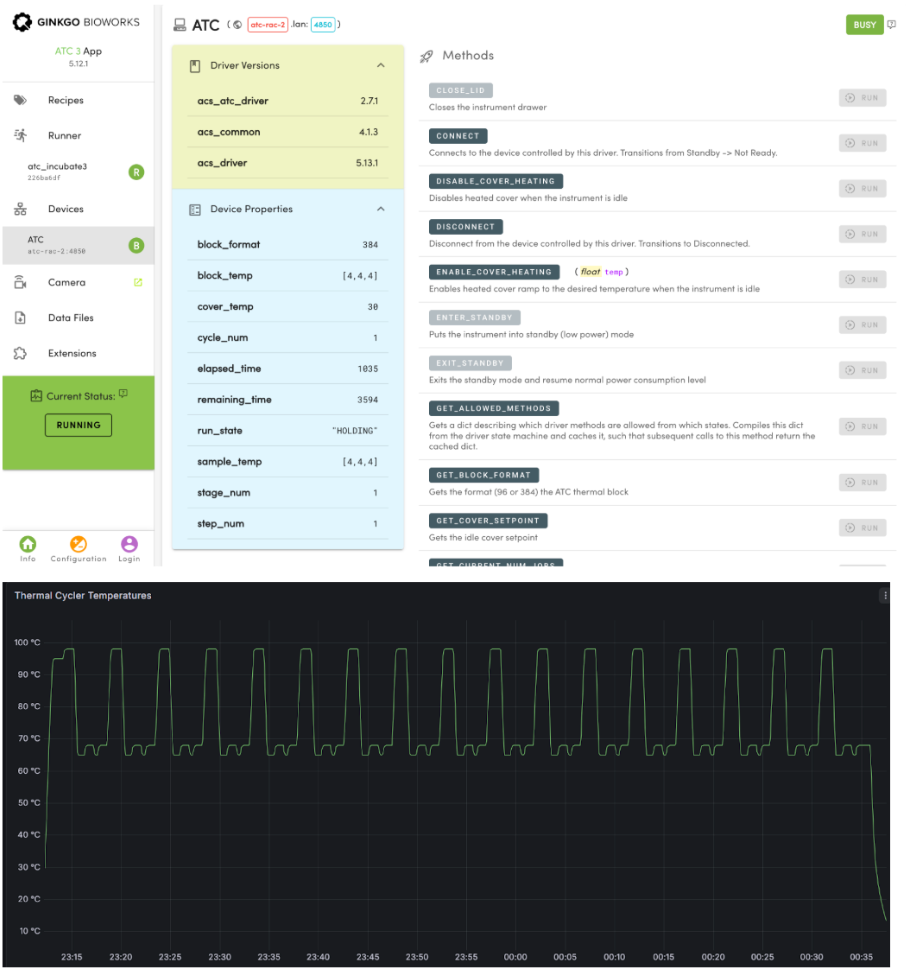
When launching the multiplexed PCR protocol, Ginkgo’s Catalyst software enabled easy customization of thermal cycling profiles, necessary for ensuring uniform representation of all required DNA oligos in the PCR reactions. Real-time temperature profiles were viewable via the ATC RAC web app and were saved in the Ginkgo Catalyst time-series database, for retrospective review (see Figure 1).
- Magnetic bead-based PCR amplicon cleanup: To purify the output PCR amplicons before the next processing step, magnetic bead-based cleanup was performed on the 384-well format Bravo liquid handling RAC as well. This protocol step can be repeated following any of the downstream enzymatic reactions, to purify DNA outputs as desired. Since magnetic bead-based cleanup is broadly applicable to different input sample types (linear, circular DNA / RNA and proteins), we leveraged Ginkgo’s Catalyst software ability to parametrize key protocol inputs (number of wash cycles, elution volume etc.), to ultimately build a protocol that can be easily reused in the future.
- Enzymatic treatments: Additional enzymatic treatments were performed to increase the likelihood of obtaining the desired full length DNA sequences downstream. Similarly to the multiplexed PCR protocol, the Bravo and ATC RACs were used, with Ginkgo’s Catalyst software enabling the thermal cycling profile customization.
- DNA digestion and ligation reactions: Same logic was followed when performing the downstream multi-piece DNA assemblies into longer gene-length fragments.
- Gene-length PCR: “Finishing” PCR reactions were used to amplify the desired full length DNA products.
Results
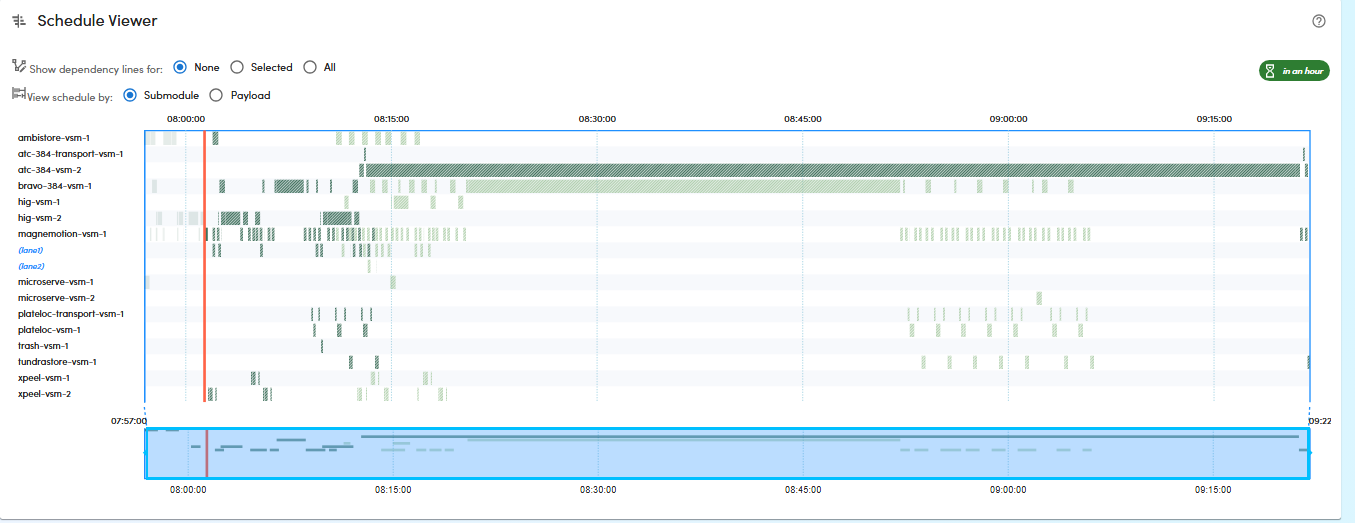
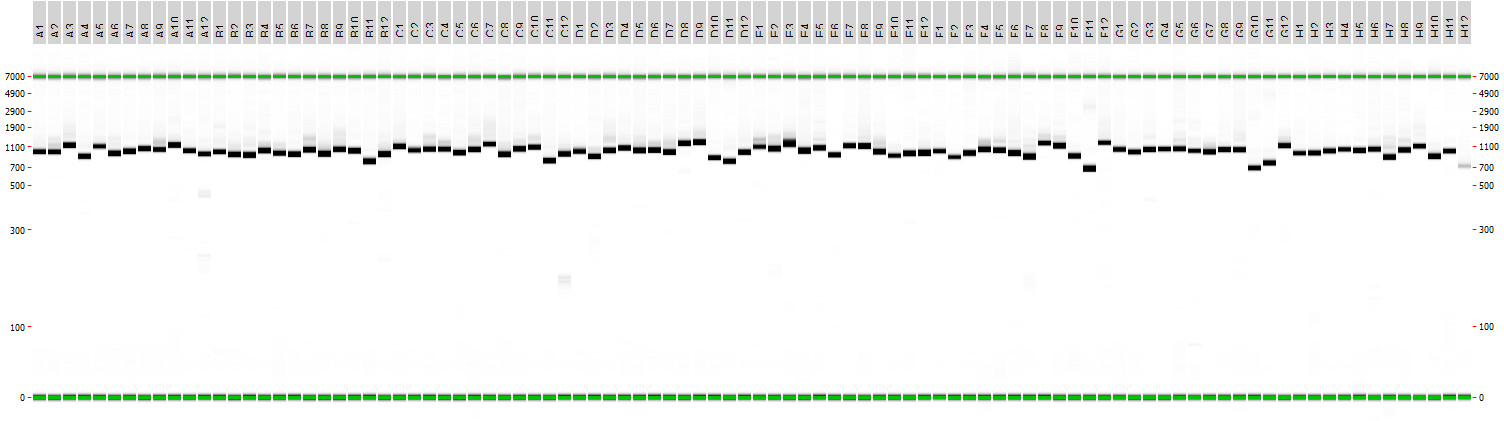
96 gene-sized fragments of unique length and sequence were successfully synthesized, using the above protocol steps (see Figure 2), as shown by the offline capillary electrophoresis QC results (see Figure 3 and 4). No contaminating DNA byproducts were observed, proving high-fidelity of our gene synthesis workflow.

Conclusions
This technical note demonstrates how the RAC platform can be successfully leveraged to execute complex workflows, here gene synthesis, yielding high-quality results.
Complex, multi-step workflows can be easily set up via the protocol chaining functionality of Ginkgo’s Catalyst software, and key individual protocols, here magnetic bead-based cleanup, can be easily reused, thanks to the built-in parametrization features. Multiple, different protocol runs can be interleaved to increase throughput, while still ensuring high-quality results and end-to-end traceability via key metadata capture (e.g. thermal cycling profile metadata).
In the future, further walkaway time gains can be achieved via a simple capillary electrophoresis RAC addition, and further conditional chaining, based on upstream workflow QC results, with downstream workflows, using newly built gene-sized fragments.
References
- Galanie, S. et al. (2015) ‘Complete biosynthesis of opioids in yeast’, Science, 349(6252), pp. 1095–1100.
- Cao, J. et al. (2021) ‘High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies’, Nature Communications, 12(4138).
- Wu, S. et al. (2024) ‘High titer expression of antibodies using linear expression cassettes for early-stage functional screening’, Protein Engineering, Design and Selection, 37, gzae012.
- Eroshenko, N. et al. (2012) ‘Gene Assembly from Chip-Synthesized Oligonucleotides’, Curr. Protoc. Chem. Biol. 2012, ch110190.
- Engler, C. et al. (2008) ‘A one pot, one step, precision cloning method with high throughput capability’, PloS One 3, e3647.
{{data}}
Subscribe to receive future tech notes
- 96 Genes synthesized with high fidelity
- 0 Runtime errors
- >8 hours Of walkaway time gained
Real Labs. Real Impact.
Learn more about our technology notes.


